Group Leader/s
intro
Understanding the molecular mechanisms that determine virus-host compatibility and disease is a major driving force for biologists and plant pathologists. Bearing this idea in mind, we investigate the crosstalk between virus infection and plant innate immunity. Thus, while normal, coevolved interactions of viruses with the host plant are optimally balanced such that infection is achieved without causing damage to the host, viruses may cause dangerous outbreaks and disease if this subtle equilibrium is disturbed.
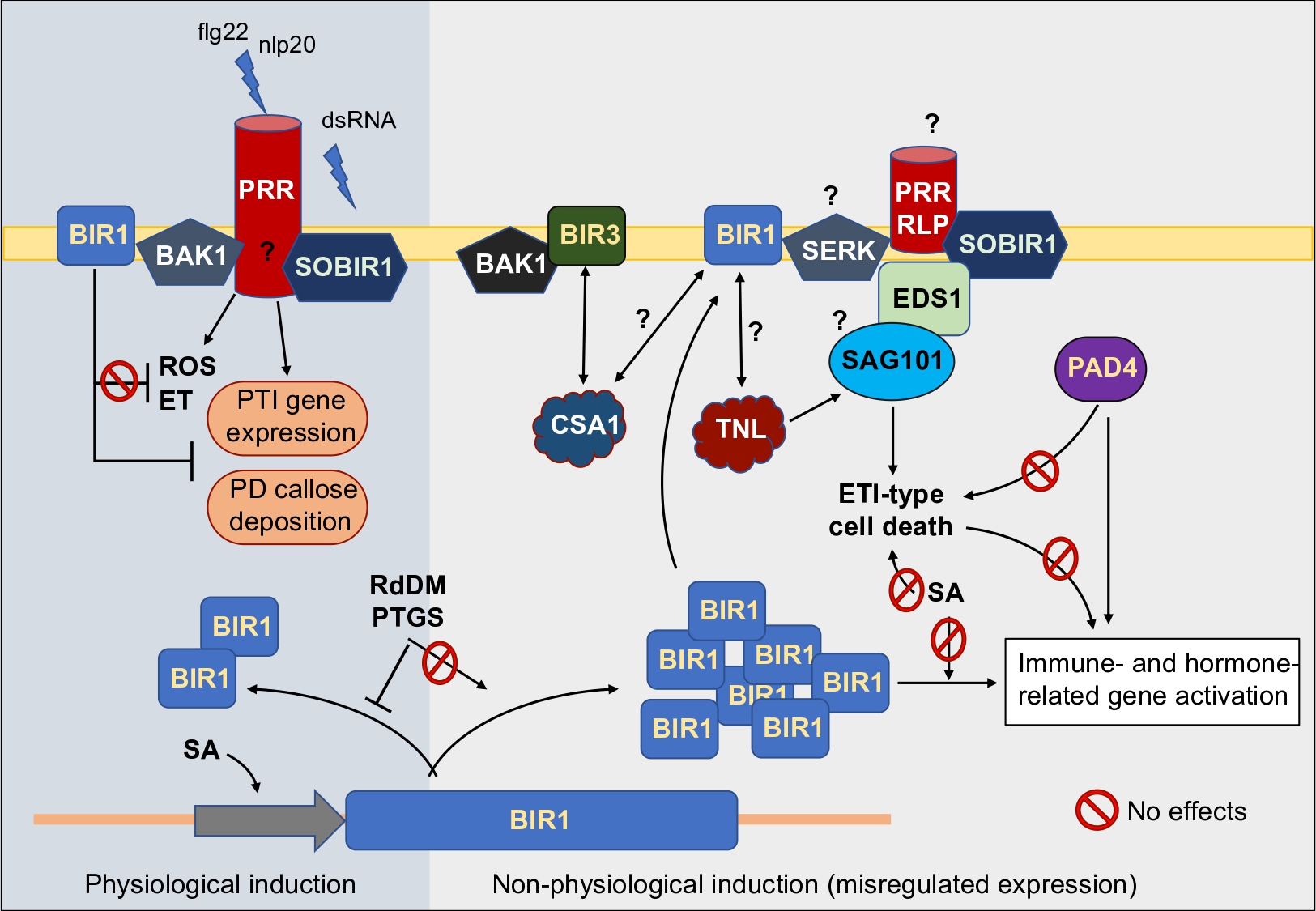
Plant immunity allows to perceive non-self and mount responses to keep microbes in check. Growing evidence indicates that viruses elicit immune responses, which may contribute to combat viral infections in plants. We used reverse genetics tools to study the role of multiple immune (co)receptor and immune regulators in this antiviral response. Among them, we identified the cell-surface receptor-like kinase BIR1 as a negative regulator of antiviral immunity. Virus infection stimulates the accumulation of SA, which in turn triggers transcriptional activation of BIR1. Since BIR1 is an essential component for immune attenuation in plants, our works has been focused to understand the mechanisms of BIR1 homeostasis. We found that DNA methylation constitutively regulates BIR1 transcription, while post-transcriptional RNA silencing removes the excess of BIR1 transcripts when BIR1 is transcriptionally activated, demonstrating that RNA silencing is a regulatory checkpoint of the immune response. We have further explored the consequences of BIR1 induction. BIR1 induction at physiological levels interferes with both the expression pattern-triggered immunity (PTI)-related genes and deposition of callose at the plasmodesmata triggered by immune elicitors. BIR1 induction may be a pathogen-driven mechanism to modulate plant defenses during infections, helping to maintain host-pathogen equilibrium. We found that non-physiological expression of BIR1 results in severe growth defects and cell-death and a broad misregulation of immunity. In recent years, we also put the focus in deciphering novel components of the BIR1-dependent antiviral signaling and how they function.
We identified several master regulators of PTI and effector triggered immunity (ETI) signaling cascades, including EDS1 and SOBIR1, that are required for the ETI-like of cell death phenotypes observed when BIR1 is expressed at non-physiological levels. We hypothesize that BIR1, previously thought to represent a negative regulator of plant immunity, is a guarded protein surveilled by unknown TNL-type resistant proteins. Thus, TNLs may monitor the integrity of BIR1 or BIR1-co-receptor complexes, with SOBIR1 working alongside EDS1 to transduce signals downstream of TNL.
Recently, we set up an experimental pipeline to search for novel BIR1-interacting protein candidates in infected plants as well as BIR1-responsive genes. Also of interest, we study how plant viruses are perceived by extracellular immune receptors and how the perception of plant viruses connects to the downstream immune signaling pathways.
BIR1, due to its role as a repressor of basal- and effector-triggered immunity, is a promising tool for engineering resistance against pathogen infections in plants. Since future sustainable agriculture should rely on knowledge-based approaches that make use of an in-depth understanding of plant-virus interactions, our results are expected to contribute to the development of sustained crop production that uses improved plant cultivars avoiding treatments with environmentally toxic chemicals. Our research is oriented toward the challenge identified as “Bioeconomía: sostenibilidad de los sistemas de producción primaria y forestales, seguridad y calidad alimentaria, investigación marina y marítima y bioproductos”.

Members
| César Llave Correas |
| Malgorzata Ciska |
| Carmen Robinson Pastor |
| Ana Yepes Melchor |

Selected Publications
Guzmán-Benito I, Robinson C, Hua C, Sede A, Elvira-González L, Punzón I, Heinlein M, Nürnberger T, LLAVE C*. [2023]. The receptor-like kinase BIR1 inhibits elicitor-induced plasmodesmata callose deposition and PTI gene expression and requires EDS1 and SOBIR1 to cause dose-dependent cell-death in Arabidopsis. BioRxiv doi:10.1101/2023.06.23.546234
Pitzalis N , Amari K , Graindorge S, Pflieger D , Donaire L, Wassenegger M, Llave C*, M. Heinlein* [2020]. Turnip mosaic virus in oilseed rape activates networks of sRNA-mediated interactions between viral and host genomes. Communications Biology doi: 10.1038/s42003-020-01427-y
Guzmán-Benito I, Donaire L, Amorim-Silva V, Vallarino JG, Esteban A, Wierzbicki AT, Ruiz-Ferrer V, Llave C* [2019]. The immune repressor BIR1 contributes to antiviral defense and undergoes transcriptional and post-transcriptional regulation during viral infections. New Phytologist 224: 421-438. doi: 10.1111/nph.15931. [Epub ahead of print]
Diezma-Navas L, Pérez-González A, Artaza H, Alonso L, Caro E, Llave C*, Ruiz-Ferrer V* [2019]. Crosstalk between epigenetic silencing and infection by Tobacco rattle virus in Arabidopsis. Molecular Plant Pathology 20: 1439-1452. DOI: 10.1111/mpp.12850
Donaire L, Llave C* [2019]. Computational Workflow for Small RNA Profiling in Virus-Infected Plants. Methods Mol Biol. 2019;2028:185-214. doi: 10.1007/978-1-4939-9635-3_11.
Fernández-Calvino L, Osorio S, Hernández ML, Hamada IB, Del Toro FJ, Donaire L, Yu A, Bustos R, Fernie AR, Martínez-Rivas JM and Llave C* [2014]. Virus-Induced Alterations in Primary Metabolism Modulate Susceptibility to Tobacco rattle virus in Arabidopsis. Plant Physiology 166(4):1821-38. doi: 10.1104/pp.114.250340.
Fernández-Calvino L, Guzmán-Benito I, Del Toro FJ, Donaire L, Castro-Sanz AB, Ruíz-Ferrer V, Llave C* [2016]. Activation of senescence-associated dark-inducible genes during infection contributes to enhance susceptibility to plant viruses. Molecular Plant Pathololy 17(1):3-15. doi: 10.1111/mpp.12257.
Llave C [2016]. Dynamic cross-talk between host primary metabolism and viruses during infections in plants. Current Opinion in Virology 19, 50-55. doi: 10.1016/j.coviro.2016.06.013
Fernández-Calvino L, Martínez-Priego L, Szabo EZ, Guzmán-Benito I, González I, Canto T, Lakatos L, Llave C* [2016]. Tobacco rattle virus 16K silencing suppressor binds AGO4 and inhibits formation of RNA silencing complexes. Journal of General Virology 97(1):246-57. doi: 10.1099/jgv.0.000323.
Llave, C., Xie, Z., Kasschau, K.D., and Carrington, J.C. [2002]. Cleavage of Scarecrow-like mRNA targets direct by a class of Arabidosis miRNA. Science 297, 2053-2056.
Llave, C., Kasschau, K.D., Rector, M.A. and Carrington, J.C. [2002]. Endogenous and silencing-associated small RNAs in plants. Plant Cell 14, 1605-1619.
Donaire L, Barajas D, Martínez-García B, Martínez-Priego L, Pagán I, Llave C* [2008]. Structural and genetic requirements for the biogenesis of tobacco rattle virus-derived small interfering RNAs. Journal of Virology 82(11):5167-77
Funding
1. Biodiversity and functional genomics of endogenous small RNAs induced upon biotic and abiotic stresses in plants (plasmar). Ref: GEN2003-20222-CO2-00. MCYT / GENOPLANTE / GABI. 2004-2006. 204.700 €. César Llave (PI)
2. Genómica funcional de micro-ARNs y otros pequeños ARNs como represores de la expresión de genomas virales. Ref. GR/SAL/0831/2004.Comunidad de Madrid. 2005. 43.500 €. César Llave (PI)
3. Análisis de la estructura y función de microARNs y otros pequeños ARNs como reguladores génicos durante las infecciones virales de las plantas. Ref. BIO2006-13107. MEC. 2006-2009. : 212.960 €. César Llave (PI)
4. Identificación de factores y mecanismos celulares implicados en interacciones compatibles entre virus y plantas. Ref. CCG07-CSIC/GEN-1804.Comunidad de Madrid. 2008. 27.200 €. César Llave (PI)
5. Generación de herramientas genómicas en olivo y su aplicación en el análisis de la calidad de fruto y del aceite, y de caracteres agronómicos. Proyecto OLEAGEN. Fundación GENOMA ESPAÑA. 2008-2011. 97.718 €. César Llave (PI at CIB)
6. Análisis funcional de la actividad reguladorade pequeñós RNAs virales sobre los genomas del virus y del huésped e implicaciones en el desarrollo de infecciones virales en plantas. Ref. BIO2009-12004. MICINN. 2010-2012. César Llave (IP). 172.000€
7. Red Nacional de Virología de Plantas. Ref. BIO2009-07108-E. MICINN. 2010. César Llave (IP). 25.000 €
8. Functional significance of microRNA-regulatory nodes in plant-virus interactions: Comparative miRNA profiling and functional degradome. Ref. 2011BR0078. MINECO-CSIC. 2013-2014. César Llave (IP). 30.000 €
9. Functional analysis of virus-inducible, RNA silencing-associated regulatory nodes and evolution of epigenetic marks associated to plant-virus interactions. Ref. BIO2012-39973. Ministerio de Economía y Competitividad. 2013-2015. César Llave (IP). 130.000 €
10. Genes and mutants affecting viral infections in rapeseed (GAMAVIR). Ref. PCIN-2013-064. Ministerio de Economia y Competitividad. 2013-2015. 100.000€
11. Cross-talk interactions between small RNA-directed RNA silencing and basal innate immunity during viral infections in plants. Ref. BIO2015-70752. MINECO. 2016-2018. 130.000€
12. Characterization of the immune response during viral infections in plants: Regulation and signaling pathways dependent on the immune repressor BIR1. Ref: RTI2018-096979-B-I00. MINECO. 2019-2021. 133.100€
13. Novel immune signaling pathways associated to antiviral responses in plants. Ref. LINKA20415. Programa i-LINK 2021. MICINN. 2022-2023. César Llave (IP). 20.066,68€.
14. Novel immune signaling pathways dependent on the plant immune repressor BIR1. Ref. PID2021-127982NB-I00. MICINN. 2022-2024. César Llave (IP). 134.000€.
15. Searching for viral elicitors of the immune response in plants: Characterization of the virus-associated secretome. Ref: TED2021-131408B-I00. MICINN. 2023-2024. César Llave (IP). 130.000€

